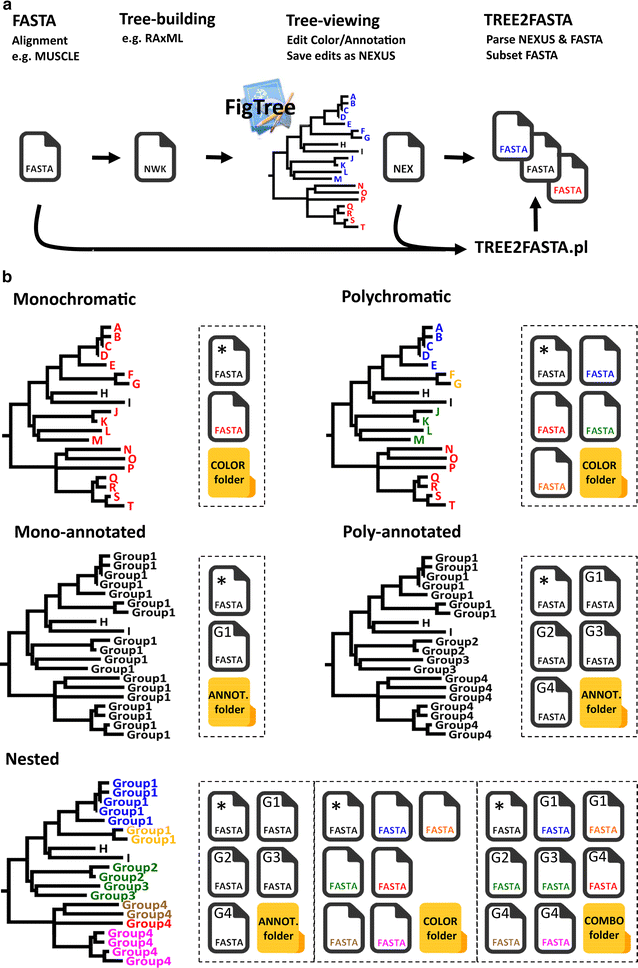
TREE2FASTA: a flexible Perl script for batch extraction of FASTA sequences from exploratory phylogenetic trees | BMC Research Notes | Full Text

PDF) Risku & Kanervio: Doctoral and Regular Research on Principals in Finland 2000 - 2010 | Pekka Kanervio - Academia.edu

Can anyone tell me how to convert Fasta format (.fasta) multiple sequences into (.nhx OR nh) format for phylogenetic predictions? | ResearchGate